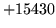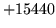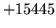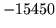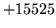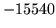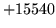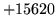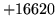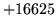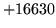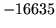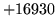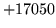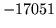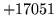Summary of Errors by Tool and Module
- HCOPY
-
-
- Non-existent part of file specified
HCOPY needed to access a non-existent part of the input file.
Check that the times are specified correctly, that the label file
contains enough labels and that it corresponds to the data file.
-
- Label file formatted incorrectly
HCOPY is only able to properly copy label files with the
same number of levels/alternatives. When using labels with multiple
alternatives only the first one is used to determine segment
boundaries.
-
- Appending files of different type/size/rate
Files that are joined together must have the same parameter kind and
sample rate.
-
- ALIEN format set
Input/output format has been set to ALIEN, ensure that
this was intended.
- HLIST
-
- HLED
-
-
- Edit script syntax error
The HLED command script contains a syntax error, check the
input script against the descriptions of each command in
section 17.10 or obtained by running HLEd -Q.
-
- Operation invalid
You have either exceeded HLED limits on the number of
boundaries that can be specified, tried to perform an operation on
a non-existent level or tried to sort an auxiliary level into time
order. None of these operations are supported.
-
- Cannot find pronunciation
The dictionary does not contain a valid pronunciation (only occurs
when attempting expansion from a dictionary).
-
- ALIEN format set
Input/output format has been set to ALIEN, ensure that
this was intended.
- HLSTATS
-
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- No operation specified
You have invoked HLSTATS but have not specified an operation
to be performed.
-
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HDMAN
-
-
- Limit exceeded
HDMAN has several built in limits on the number of different
pronunciation, phones, contexts and command arguments. This error
occurs when you try to exceed one of them.
-
- Item not found
Could not find item for deletion. Check that it actually occurs in the
dictionary.
-
- Edit script file syntax error
The HDMAN command script contains a syntax error, check the
input script against the descriptions of each command in
section 17.5 or obtained by running HDMan -Q.
-
- Dictionary file syntax error
One of the input dictionaries contained a syntax error. Ensure that
it is in a HTK readable form (see section 12.7).
-
- Word out of order in dictionary error
Entries in the dictionary must be sorted into alphabetical (ASCII) order.
- HSLAB
-
-
- ALIEN format set
Input/output format has been set to ALIEN, ensure that
this was intended.
- HCOMPV
-
-
- HMM does not appear in HMMSet
Supplied HMM filename does not appear in HMMSet. Check
correspondence between HMM filename and HMMSet.
-
- Not enough data to calculate variance
There are not enough frames of data to evaluate a
reliable estimate of variance. Use more data.
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- Needs continuous models
HCompV can only operate on models with an HMM set kind
of PLAINHS or SHAREDHS.
-
- Speaker pattern matching failure
The specified speaker pattern could not be matched against a
given utterance file name.
-
- Data does not match HMM
An aspect of the data does not match the equivalent aspect in
the HMMSet. Check the parameter kind of the data.
-
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HINIT
-
-
- Unknown update flag
Unknown flag set by -u option, use combinations of
tmvw.
-
- Too little data
Not enough data to reliably estimate parameters. Use more
training data.
-
- Segment with fewer frames than model states
Segment may be too short to be matched to model, do not use
this segment for training.
-
- Cannot mix covariance kind in a single mix
Covariance kind of all mixture components in any one state must be
the same.
-
- Bad covariance kind
Covariance kind of mixture component must be either
FULLC or DIAGC.
-
- No best mix found
The Viterbi mixture component allocation failed to find a most likely
component with this data. Check that data is not corrupt and
that parameter values produced by the initial uniform segmentation
are reasonable.
-
- No path through segment
The Viterbi segmentation failed to find a path through
model with this data. Check that data is not corrupt and that
a valid path exists through the model.
-
- Zero occurrence count
Parameter has had no data assigned to it and cannot be
updated. Ensure that each parameter can be estimated by
using more training data or fewer parameters.
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- HMM not found
HMM missing from HMMSet. Check that the HMMSet is complete and
has not been corrupted.
-
- Data does not match HMM
An aspect of the data does not match the equivalent aspect in
the HMMSet. Check the parameter kind of the data.
-
- Index out of range
Trying to access a mixture component or VQ index beyond the
range in the current HMM.
-
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HREST
-
-
- Unknown update flag
Unknown flag set by -u option, use combinations of
tmvw.
-
- Too few training examples
There are fewer training examples than the minimum set by the
-m option (default 3). Either reduce the value specified
by -m or use more training examples.
-
- Zero occurrence count
Parameter has had no data assigned to it and cannot be
updated. Ensure that each parameter can be estimated by
using more training data or fewer parameters.
-
- Floor too high
Mix weight floor has been set so high that the sum over all
mixture components exceeds unity. Reduce the floor value.
-
- Defunct Mix X.Y.Z
Not enough training data to re-estimate the covariance vector of
mixture component Z in stream Y of state X. The weight of the mixture
component is set to 0.0 and it will never recover even with further
training.
-
- No training data
None of the supplied training data could be used to
re-estimate the model. Data may be corrupt or has been floored.
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- Data does not match HMM
An aspect of the data does not match the equivalent aspect in
the HMMSet. Check the parameter kind of the data.
-
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HEREST
-
-
- Unknown update flag
Unknown flag set by -u option, use combinations of
tmvw.
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- No transitions
No transition out of an emitting state, ensure that
there is a transition path from beginning to end of model.
-
- Floor too high
Mix weight floor has been set so high that the sum over all
mixture components exceeds unity. Reduce the floor value.
-
- No mixtures above floor
None of the mixture component weights are greater than the floor value,
reduce the floor value.
-
- Zero occurrence count
Parameter has had no data assigned to it and cannot be
updated. Ensure that each parameter can be estimated by
using more training data or fewer parameters.
-
- Not enough training examples
Model was not updated as there were not enough training examples.
Either reduce the minimum specified by -m or
use more data.
-
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HSMOOTH
-
-
- Unknown update flag
Unknown flag set by -u option, use combinations of
tmvw.
-
- Invalid HMM set kind
HSMOOTH can only be used if HMM set kind is either
DISCRETE or TIED.
-
- Too many monophones in list
HSMOOTH is limited to HMMSets containing fewer than
500 monophones.
-
- Different number of states for smoothing
Monophones and context-dependent models have differing
numbers of states.
-
- No transitions
No transition out of an emitting state, ensure that
there is a transition path from beginning to end of model.
-
- Floor too high
Mix weight floor has been set so high that the sum over all
mixture components exceeds unity. Reduce the floor value.
-
- Zero occurrence count
Parameter has had no data assigned to it and cannot be
updated. Ensure that each parameter can be estimated by
using more training data or fewer parameters.
-
- Not enough training examples
Model was not updated as there were not enough training examples.
Either reduce the minimum specified by -m or
use more data.
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
- HQUANT
-
-
- Stream widths invalid
The chosen stream widths are invalid. Check that these match the
parameter kind and are specified correctly.
-
- Data does not match codebook
Ensure that the parameter kind of the data matches that of the codebook
being generated.
- HHED
-
-
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
- Tying null or different sized items
You have executed a tie command on items which do not have the
appropriate structure or the structures are not matched. Ensure
that the item list refers only to the items that you wish to tie
together.
-
- Performing operation on no items
The item list was empty, no operation is performed.
-
- Command parameter invalid
The parameters to the command are invalid either because they
refer to parts of the model that do not exist (for instance a
state that does not appear in the model) or because they
do not represent an acceptable value (for instance HMMSet kind
is not PLAINHS, SHAREDHS, TIEDHS or
DISCRETEHS).
-
- Join parameters invalid or not set
Make sure than the join parameters (set by the JO command)
are reasonable. In particular take care that the floor is low enough
to ensure that when summed over all the mixture
components the sum is below 1.0.
-
- Cannot find matching item
Search for specified item was unsuccessful. When this occurs with
the CL or MT commands ensure that the appropriate
monophone/biphone models are in the current HMMSet.
-
- Small gConst
A small gConst indicates a very low variance in that particular
Gaussian. This could be indicative of over-training of the models.
-
- No typical state
When tying states together a search is performed for the distribution
with largest variance and all tied states share this distribution. If
this cannot be found the first in the list will be used instead.
-
- Long macro name
In general macro names should not exceed 20 characters in length.
-
- Not implemented
You have asked HHED to perform a function that is not
implemented.
-
- Invalid stream split
The specified number/width of the streams does not agree with the
parameter kind/vector size of the models.
-
- Edit script syntax error
The HHED command script contains a syntax error, check the
input script against the descriptions of each command in
section 17.8 or obtained by running HHEd -Q.
-
- Command range error
The value specified in the command script is out of range. Ensure that
the specified state exists and the the value given is valid.
-
- Stats file load error
Either loading occupation statistics for the second time or executing
an operation that needs the statistics loaded without loading them.
-
- Trees file syntax error
The trees file format did not correspond to that expected. Ensure that
the file is complete and has not been corrupted.
-
- Trees file macro/question not recognised
The question or macro referred to does not exist. Ensure that the file
is complete and has not been corrupted.
-
- Trying to synthesize for unknown model
There is no tree or prototype model for the new context. Ensure that a
tree has been constructed for the base phone.
-
- Invalid types to tree cluster
Tree clustering will only work for single Gaussian diagonal
covariance untied models of similar topology.
- HBUILD
-
-
- Mismatch between command line and language model
Ensure that the !ENTER and !EXIT words are correctly
defined and that the supplied files are of the appropriate type.
-
- Unknown word
Ensure that the word list corresponds to the language model/lattice
supplied.
- HPARSE
-
-
- Variable not defined
You have referenced a network that has not yet been defined. Check
that all networks are defined before they are referenced.
-
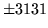
- Loop or word expansion error
There is either a mismatch between the WD_BEGIN WD_END
pairs or a triphone loop is badly formed.
-
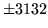
- Dictionary error
When generating a dictionary a word exceeded the maximum number of
phones, a word occurred twice or no dictionary was produced.
-
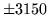
- Syntax error in HParse file
The HPARSE network definition contains a syntax error, check
the input file against the network description in
section 17.16.
- HVITE
-
-
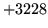
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
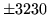
- Unsupported operation
HVITE is not able to perform the operation requested
-
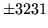
- Data does not match HMMs
There is a mismatch between the data file and the HMMSet. Ensure that
the data is parameterised in the correct format and the configuration
parameters match those used during training.
-
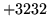
- MMF Load Error
The HMMSet does not contain a well-formed regression class tree.
-
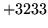
- Transcription empty
In alignment mode a segment had an empty transcription and no
boundary word was specified.
-
- ALIEN format set
Input/output format has been set to ALIEN, ensure that
this was intended.
- HRESULTS
-
-
- Empty file
The file was empty and will be skipped.
-
- Unknown label
The label did not appear in the list supplied to HResults.
This error will only occur if calculating confusion matrices so
normally the contents of the word list file will have no effect
on results.
-
- Too many labels
HRESULTS will only generate confusion statistics for a small
number of labels.
-
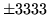
- Cannot calculate word spot results
When calculating word spotting results the label files need to have
both times and scores present.
-
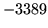
- ALIEN format set
Input format has been set to ALIEN, ensure that this was
intended.
- HSGEN
-
-
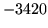
- Network malformed
The word network is malformed. The information in a node (word
and following arcs) is set incorrectly.
- HLRESCORE
-
-
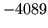
- ALIEN format set
Input/output format has been set to ALIEN, ensure that
this was intended.
- HSHELL
-
-
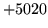
- Command line processing error
-
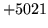
- Command line argument type error
-
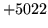
- Command line argument range error
The command line is badly formed. Ensure that it matches the
syntax and values expected by the command (check the manual
page or the syntax obtained by running HTOOL without any
arguments).
-
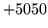
- Configuration file format error
HSHELL was unable to parse the configuration. Check that
it is of the format described in section 4.3.
-
- Script file format error
Check that the script file is just a list of file names and that
if any file names are quoted that the quotes occur in pairs.
-
- Module version syntax error
A module registered with HShell with an incorrectly formatted
version string (which should be of the form
"!HVER!HModule: Vers.str [WHO DD/MM/YY]").
-
- Too many configuration parameters
The size of the buffer used by one of the tools or modules to read
its configuration parameters was exceeded. Either reduce the total
number of configuration parameters in the file or make more of then
specific to their particular module rather than global.
-
- Configuration parameter of wrong type
The configuration parameter is of the wrong type. Check that its type
agrees with that shown in chapter 18.
-
- Configuration parameter out of range
The configuration parameter is out of range.
- HMEM
-
-
- Heap parameters invalid
You have tried to create a heap with unreasonable parameters. Adjust
these so that the growth factor is positive and the initial block
size is no larger than the maximum. For MSTAK the element
size should be 1.
-
- Heap not found
The specified heap could not be found, ensure that it has not been
deleted or memory overwritten.
-
- Heap does not support operation
The heap is of the wrong type to support the requested operation. In
particular it is not possible to Reset or Delete a
CHEAP.
-
- Wrong element size for MHEAP
You have tried to allocate an item of the wrong size from a
MHEAP. All items on a MHEAP must be of the same size.
-
- Heap not initialised
You have tried to allocate an item on a heap that has not yet been
created. Ensure that CreateHeap is called to initialise
the heap before any items are allocated from it.
-
- Freeing unseen item
You have tried to free an item from the wrong heap. This can occur
if the wrong heap is specified, the item pointer has been corrupted
or the item has already been freed implicitly by a
Reset/DeleteHeap call.
- HMATH
-
-
- Singular covariance matrix
The covariance matrix was not invertible. This may indicate a lack
of training data or linearly dependent parameters.
-
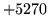
- Size mismatch
The input parameters were of incompatible sizes.
-
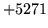
- Log of negative
Result would be logarithm of a negative number.
- HSIGP
-
-
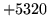
- No results for WaveToLPC
Call did not include Vectors for the results.
-
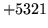
- Vector size mismatch
Input vectors were of mismatched sizes.
-
- Clamped samples during zero mean
During a zero mean operation samples were clipped as they were outside
the allowable range.
- HAUDIO
-
-
- Replay buffer not active
Attempt to access a replay buffer when one was not attached.
-
- Cannot StartAudio without measuring silence
An attempt was made to start audio input through the silence detector
without first measuring or supplying the background silence values.
-
- Audio frame size/rate invalid
The choice of frame period and window duration are invalid. Check
both these and the sample rate.
-
- Setting speech threshold below silence
The thresholds used in the speech detector have been set so that the
threshold for detecting speech is set below that of detecting silence.
- HVQ
-
-
- VQ file format error
The VQ file was incorrectly formatted. Ensure that the file is
complete and has not been corrupted.
-
- VQ file range error
A value from the VQ file was out of range. Ensure that the file is
complete and has not been corrupted.
-
- Magic number mismatch
The VQ magic number (normally based on parameter kind) does not match
that expected. Check that the parameter kind used to quantise the data
and create the VQ table matches the current parameter kind.
-
- VQ table already exists
All VQ tables must have distinct names. This error will occur if you
try to create or load a VQ table with the same name as one already
loaded.
-
- Invalid covariance kind
Entries in VQ tables must have either NULLC, FULLC or
INVDIAGC covariance kind.
-
- Node not in table
A node was missing from the VQ table. Ensure that the VQ table was
properly created or that the file was complete.
-
- Stream codebook mismatch
The number or size of streams in the VQ table does not match that
requested.
- HWAVE
-
-
- Cannot fseek/ftell
Unless the wave file is read through a pipe fseek and ftell are
expected to work correctly so that HWAVE can calculate the
file size. If this error occurs when using an input pipe, supply
the number of samples in the file using the configuration variable
NSAMPLES.
-
- File appears to be a infinite
HWAVE cannot determine the size of the file.
-
- Config parameter not set
A necessary configuration parameter has not been set. Determine the
correct value and place this in the configuration file before
re-invoking the tool.
-
- Premature end of header
HWAVE could not read the complete file header.
-
- Header contains invalid data
HWAVE was unable to successfully parse the header. The header
is invalid, of the wrong type or be a variation that HWAVE does
not handle.
-
- Header missing essential data
The header was missing a piece of information necessary for
HWAVE to load the file. Check the processing of the
input file and re-process if necessary.
-
- Premature end of data
The file ended before all the data was read correctly. Check that the
file is complete, has not been corrupted and where necessary
NSAMPLES is set correctly.
-
- Data formated incorrectly
The data could not be decoded properly. Check that the file was
complete and processed correctly.
-
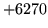
- File format invalid
The file format is not valid for the operation requested.
-
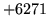
- Attempt to read outside file
You have tried to read a sample outside of the waveform file.
- HPARM
-
-
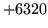
- Configuration mismatch
The data file does not match the configuration. Check the
configuration file is correct.
-
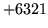
- Invalid parameter kind
Parameter kind is not valid. Check the configuration file.
-
- Conversion not possible
The specified conversion is not possible. Check the configuration is
correct and re-code the data from waveform files if necessary.
-
- Audio error
An audio error has been detected. Check the HAUDIO
configuration and the audio device.
-
- Buffer not initialised
Ensure that the buffer is used in the correct manner.
-
- Silence detection failed
The silence detector was not initialised correctly before use.
-
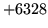
- Load/Make HMMSet failed
The model set could not be loaded due to either an error opening the
file or the data within being inconsistent.
-
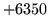
- CRC error
The CRC does not match that of the data. Check the data file is
complete and has not been corrupted.
-
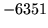
- Byte swapping not possible
HPARM will attempt to byte swap parameter files but this
may not work if the floating point representation of the machine
that generated the file is different from that which is reading it.
-
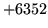
- File too short to parameterise
The file does not contain enough data to produce a single observation.
Check the file is complete and not corrupt. If it is, it should be
discarded.
-
- Unknown parameter kind
The specified parameter kind is not recognised. Refer to
section 5.18 for a list of allowable parameter kinds
and qualifiers.
-
- Invalid parameters for coding
The chosen parameters are not valid for coding. Choose different ones.
-
- Stream widths not valid
Cannot split the data into the specified number of streams. Check that
the parameter kind is correct and matches any models used.
-
- Buffer/observation mismatch
The observation parameter kind should match that of the input buffer.
Check that the configuration parameter kind is correct and matches
that of any models used.
-
- Buffer size too small for window
Calculation of delta parameters requires a window larger than the
buffer size chosen. Increase the size of the buffer.
-
- Frame not in buffer
An attempt was made to access a frame that does not appear in the
buffer. Make sure that the file actually contains the specified frame.
-
- Mean/Variance normalisation failed
The mean or variance normalisation vector from the file
specified by the normalisation dir and mask cannot be applied.
Make sure the file format is correct and the vectors are of
the right dimension.
- HLABEL
-
-
- MLF index out of range
An attempt was made to access an MLF that has not been loaded or to
load too many MLFs.
-
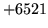
- fseek/ftell not possible
HLABEL needs random access to MLFs. This error is generated
when this is not possible (for instance if access is via a pipe).
-
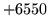
- HTK format error
-
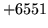
- MLF format error
-
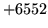
- TIMIT format error
-
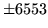
- ESPS format error
-
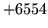
- SCRIBE format error
A label file was formatted incorrectly. Label
file formats are described in chapter 6.
-
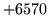
- Level out of range
Attempted to access a non-existent label level. Check that the correct
label file has been loaded.
-
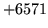
- Label out of range
Attempted to access a non-existent label. Check that the correct
label file has been loaded and that the correct level is being
accessed.
-
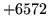
- Invalid format
The specified file format is not valid for the particular operation.
- HMODEL
-
-
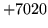
- Cannot find physical HMM
No physical HMM exists for a particular logical model. Check that the
HMMSet was loaded or created correctly.
-
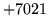
- INVDIAG internal format
Attempts to load or save models with INVDIAG covariance kind
will fail as this is a purely internal model format.
-
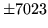
- varFloor should be variance floor
HMODEL reserves the macro name varFloorN as the
variance floor for stream N. These should be variance
macros (type v) of the correct size for the particular stream.
-
- Variance tending to 0.0
A variance has become too low. Start using a variance floor or
increase the amount of training data.
-
- Bad covariance kind
The particular functionality does not support the covariance
kind of the mixture component.
-
- HMM set incomplete or inconsistent
The HMMSet contained missing or inconsistent data. Check that the
file is complete and has not been corrupted.
-
- HMM parameters inconsistent
Some model parameters were inconsistent. Check that the file is
complete and has not been corrupted.
-
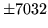
- Option mismatch
All HMMs in a particular set must have consistent options.
-
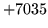
- Unknown macro
Macro does not exist. Check that the name is correct and appears
in the HMMSet.
-
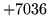
- Duplicate macro
Attempted to create a macro with the same name as one already present.
Choose a different name.
-
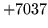
- Invalid macro
Macro had invalid type. See section 7.3 describes the
allowable macro types.
-
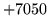
- Model file format error
-
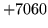
- HMM List format error
The file was formated incorrectly. Check the file is complete and
has not been corrupted.
-
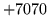
- Invalid HMM kind
Invalid HMMSet kind. Check that this is specified correctly.
-
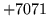
- Observation not compatible with HMMSet
Attempted to calculate an observation likelihood for an observation
not compatible with the HMMSet. Check that the parameter kind is
set correctly.
- HTRAIN
-
-
- Clustering failed
Almost certainly due to a lack of data, reduce the
number of clusters requested or increase amount of data.
-
- Accumulator file format error
Cannot read an item from an accumulator file. Check
that file is complete and not corrupted.
-
- Unsupported covariance kind
Covariance kind must be either FULLC, DIAGC or
INVDIAGC.
-
- Item out of range
Attempt made to access data beyond expected range. Check that
the item number is correct.
-
- Tree size must be power of 2
Requested codebook size must be a power of 2 when
using tree based clustering.
-
- Segment empty
Empty data segment in file. Check that file has not
become corrupted and that the start and end segment times
are correct.
- HUTIL
-
-
- HMMSet empty
A scan was initiated for a HMMSet with no members.
-
- Item list parse error
The item list syntax was incorrect. Check the item list specification
in section 17.8.
-
- Item list type error
Each item in a particular list should be of the same type and size.
-
- Stats file format error
Stats file is of wrong format. Note the format of the stats file has
changed in HTK_V2.0 and old files will need converting to the new
format.
-
- Stats file model error
A model name encountered in the stats file is invalid check that the
model set corresponds to that used to generate the stats file and that
the stats file is complete and has not been corrupted.
-
- Accessing non-existent macro
Attempt to perform operation on non-existent macro.
-
- Member id out of range
Attempt to perform set operation on out of range member.
- HFB
-
-
- Unknown model
Model in HMM List not found in HMMSet, check that the
correct HMM List is being used.
-
- Invalid output probability
Mixture component probability has not been set. This should
not occur in normal use.
-
- Beta prune failed on taper
Utterance is possibly too short for minimum duration
of model sequence. Check transcription.
-
- No path through utterance
No path was found on the beta training pass, relax the
pruning threshold.
-
- Empty label file
No labels found in label file, check label file.
-
- Single-pass retraining data mismatch
Paired training files must contain the same number of observations.
Use original data to re-parameterise.
-
- HMM with unreachable states
HMM has an unreachable state, check transition matrix.
-
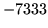
- Transition matrix with discontinuity
Check transition matrix.
-
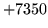
- Data does not match HMM
An aspect of the data does not match the equivalent aspect in
the HMMSet. Check the parameter kind of the data.
- HDICT
-
-
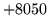
- Dictionary file format error
The dictionary file is not correctly formatted.
Section 12.7 describes the HTK dictionary file format.
- HLM
-
-
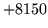
- LM syntax error
The language model file was formatted incorrectly. Check the file is
complete and has not been corrupted.
-
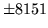
- LM range error
The specified value(s) for the language model probability are not
valid. Check the input files are correct.
- HNET
-
-
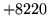
- No such word
The specified word does not exist or does not have a valid
pronunciation.
-
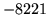
- Duplicate pronunciations removed
During network generations duplicate identical pronunciations
of the same word are removed.
-
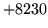
- Contexts not consistent
HNET can only deal with the standard HTK method for specifying
context left-phone+right and will only allow context free
phones if they are context independent and only form part of the word.
This may be indicative of an inconsistency between the symbols in the dictionary
and the hmms as defined. There may be a model/phone in the dictionary that has
not been defined in the HMM list or may not have a corresponding model.
See also section 12.8 on context expansion.
-
- No such model
A particular model could not be found. Make sure that the network is
being expanded in the correct fashion and then ensure that your HMM
list will cover all required contexts.
-
- Lattice badly formed
Could not convert lattice to network. The lattice should have a single
well defined start and a single well defined end. When cross word
expansion is being performed the number of !NULL words that
can be concatenated in a string is limited.
-
- Lattice format error
The lattice file is formatted incorrectly. Ensure that the lattice
is of the format described in chapter 20.
-
- Lattice file data error
The value specified in the lattice file is invalid.
-
- Lattice file with multiple start/end nodes
A lattice should have only one well defined start node and one
well defined end node.
-
- Lattice with invalid sub lattices
The sub lattices referred to by the main lattices are
malformed.
- HREC
-
-
- Invalid HMM
One of the HMMs in the network is invalid. Check that the HMMSet
has been correctly initialised.
-
- Network structure invalid
The network is incorrectly structured. Take care to avoid loops
that can be traversed without consuming observations (this may occur
if you introduce any 'tee' words in which all the models making up that
word contain tee-transitions). Also ensure that the recogniser and
the network have been created and initialised correctly.
-
- Lattice structure invalid
The lattice was incorrectly formed. Ensure that the lattice was
created properly.
-
- Recogniser not initialised correctly
Ensure the recogniser is initialised and used correctly.
-
- Data does not match HMMs
The observation does not match the HMM structure. Check the parameter
kind of the data and ensure that the data is matched to the HMMs.
- HLAT
-
-
- Lattice incompatible with dictionary
The lattice refers to a pronunciation variant (filed
v=) that doesn't exist in the current dictionary.
-
- Lattice structure invalid
The lattice does not meet the requirements for some operation.
All lattices must have unique start and end nodes and for many
operations the lattices need to be acyclic (i.e. be a
Directed Acyclic Graph).
-
- Start or end word not found
The specified lattice start or end word could not be found in
the dictionary.
-
- Lattice end node label invalid
The lattice end node must either be labelled with !NULL
or the specified end word (default: !SENT_END)
-
- LLF file not found
The specified LLF file could not be found or isn't in the
right format.
-
- Lattice not found in LLF file
A lattice couldn't be found in the LLF file. Note that the
order in the LLF file must correspond to the order of processing.
-
- Lattice not found
The specified lattice file could not be opened.
-
- Lattice operation not supported
The requested operation is not supported, yet.
-
- Lattice processing sanity check failed
During processing an internal sanity check failed. This should
never happen..
- HGRAF
-
-
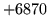
- X11 error
Ensure that the DISPLAY variable is set and that the
X11 window system is configured correctly.
- LCMAP
-
-
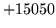
- Unlikely num map entries[n] in XYZ
A negative or infeasibly large number of class map entries
have been specified.
-
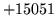
- ReadMapHeader: UNKxxx configs must be set for hdrless map
There is no header on the map so you must set UNKNOWNID and UNKNOWNNAME.
-
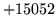
- No name in XYZ
No NAME header in class map.
-
- Unknown escmode XYZ in XYZ
ESCMODE header must specify either HTK or RAW.
-
- Class name XYZ duplicate in XYZ
Two classes in the class map have the same name, which is not allowed.
-
- Bad index n for class XYZ in XYZ
A class index less than 1 or greater than or equal to BASEWORDNDX (defined
at compile time in LWMAP - default is 65536) was found
in the class map. If you need more than BASEWORDNDX
classes then you must recompile HTK with a new base word
value.
-
- Number of entries = n for class XYZ in XYZ
There must be at least one member in each class - empty
classes are not allowed.
-
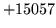
- Bad type XYZ for class XYZ in XYZ
Classes must be defined using either IN or NOTIN.
-
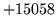
- A class is in its own exclusive list. This typically happens when a class map
is specified as a plain list of words. Such list is by default assumed to be
a list of words excluded from class !!UNK. The error is triggered when !!UNK
is in the word list. !!UNK must be removed from the list.
- LWMAP
-
-
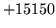
- Word list/word map file format error
Check that the word list/word map file is correctly formatted.
-
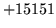
- Unlikely num map entries[n] in XYZ
A negative or infeasibly large number of word map entries
have been specified.
-
- No NAME header in XYZ
No NAME header in word map.
-
- No SEQNO header in XYZ
No SEQNO header in word map.
-
- Unknown escmode XYZ in XYZ
ESCMODE header must specify either HTK or RAW.
-
- Word name XYZ is duplicated in XYZ
There are duplicate words in the word map, which is not allowed.
- LUTIL
-
-
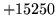
- Header format error
Ensure that word maps and/or n-gram files used by the program start
with the appropriate header.
- LGBASE
-
-
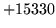
- n-gram file consistency check failure
The n-gram file is incompatible with other resources used by the
program.
-
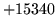
- File XYZ is n-gram but inset is n-gram
The specified input gram file is not of the expected gram size.
-
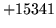
- Requested N[n] greater than gram size [n]
An n-gram was requested which was larger than any of those
supplied in the input files.
-
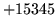
- n-grams out of order
The input gram file is not correctly sorted.
-
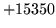
- n-gram file format error
Ensure that n-gram files used by the program are formatted correctly
and start with the appropriate header.
- LMODEL
-
-
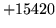
- Cannot find n-gram component
The internal structure of the language model is corrupted. This error
is usually caused when an n-gram 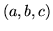 is encountered without
the presence of n-gram
is encountered without
the presence of n-gram 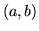 .
.
-
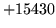
- Incompatible probability kind in conversion
The currently used language model does not allow the required
conversion operation. This error is caused by attempting to prune a
model stored in the ultra file format.
-
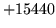
- Cannot prune models in ultra format
Pruning of language models stored in ultra file format is not
supported.
-
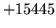
- Word ID size error
Language models with vocabularies of over 65,536 words require the
use of larger word identifiers. This is a sanity check error.
-
- Word XYZ not in unigrams - skipping n-gram.
There should be a unigram count for each word in other length grams.
-
- Language model file format error
The language model file is formatted incorrectly. Check the file is
complete and has not been corrupted.
-
- Extraneous line warning
Extra lines were found on the end of a file and are being ignored.
-
- Model order reduced
Due to the effects of pruning the model order is automatically reduced.
- LPCALC
-
-
- Unable to find FLEntry to attach
Indicates that the LM data structures are corrupt. This is normally caused
by NGram files which have not been sorted.
-
- Attempt to overwrite entries when attaching
Indicates that the LM structure is corrupt. Ensure that the word map file
used is suitable for decoding the NGram database files.
-
- n-gram cutoff out of range
An inapplicable cutoff was ignored.
-
- Pruning error
The pruning parameters specified are not compatible with the parameters
of the language model.
- LPMERGE
-
-
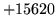
- Unable to find word in any model
Indicates that the target model vocabulary contains a word which cannot
be found in any of the source models.
- LPLEX
-
-
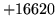
- symbol XYZ not in word list
The sentence start symbol, sentence end symbol and OOV symbol (only if
OOVs are to be included in the perplexity calculation) must be in the
language model's vocabulary. Note that the vocabulary list is either
specified with the -w option or is implicitly derived from the
language model.
-
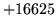
- Unable to find word XYZ in any model
Ensure that all words in the vocabulary list specified with the -w
option are present in at least one of the language models.
-
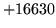
- Maximum number of unique OOVs reached
Too many OOVs encountered in the input text.
-
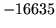
- Transcription file fn is empty
The label file does not contain any words.
-
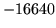
- Word too long, will be split: XYZ
The word read from the input stream is of over 200 characters.
-
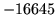
- Text buffer size exceeded (n)
The maximum number of words allowed in a single utterance has been
reached.
-
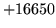
- Maximum utterance length in a label file exceeded (limit
is compiled to be n tokens)
No label file utterance end has been encountered within
n tokens - perhaps this is a text file and you forgot
to pass the -t option?
- HLMCOPY
-
-
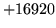
- Maximum number of phones reached
When HLMCOPY is used to copy dictionaries, the target dictionary's
phone table is composed by combining the phone tables of all source
dictionaries. Check that the number of different phones resulting from
combining the phone tables of the source dictionaries does not exceed the
internally set limit.
-
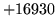
- Cannot find definition for word XYZ
When copying dictionaries, ensure that each word in the vocabulary list
occurs in at least one source dictionary.
- CLUSTER
-
-
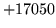
- Word XYZ found in class map but not in word map
All words in the class map must be found in the word map too.
-
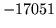
- Unknown word token XYZ was explicitly given with -u, but
does not occur in the word map
This warning appears if you specify an unknown word token
which is not found in the word map.
-
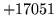
- Token not found in word list
Sentence start, end and unknown (if used) tokens must be found
in the word map.
-
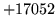
- Not all words were assigned to classes
A classmap was imported which did not include all words in the
word map.
-
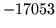
- Word XYZ is in word map but not in any gram files
The stated word will remain in whichever class it is already
in - either as defaulted to or supplied via the input class map.
Back to HTK site
See front page for HTK Authors
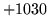
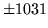
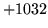
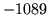
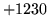
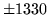
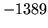
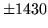
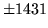
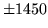
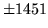
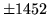
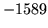
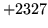
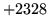
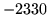
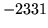
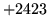
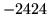
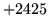
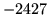
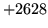
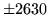
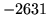
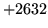
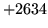
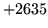
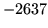
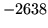
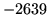
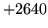
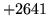
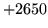
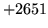
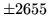
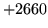
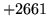
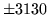
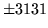
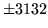
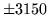
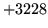
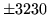
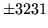
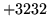
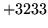
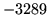
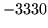
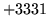
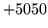
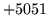
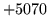
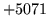
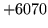
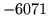
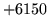
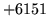
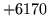
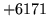
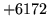
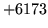
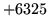
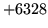
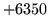
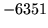
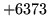
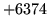
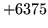
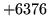
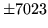
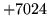
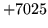
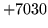
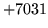
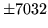
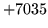
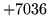
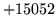
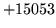
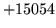
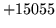
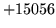
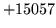
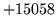
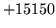
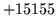
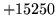
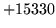
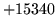
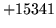
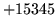
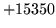
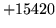
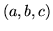 is encountered without
the presence of n-gram
is encountered without
the presence of n-gram 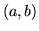 .
.